Pathways /
Chromatin remodeling/DNA methylation
Overview
The process of chromatin remodeling promotes the coordinated adjustment of chromatin structure to facilitate access to condensed regions of DNA for activation of gene transcription and control of gene expression. Chromatin remodeling is achieved through histone modifications such as DNA methylation, an epigenetic control mechanism that switches genes on and off. Chromatin remodeling may be activated or inhibited by DNA methylation and is carried out by histone methyltransferases. [1]
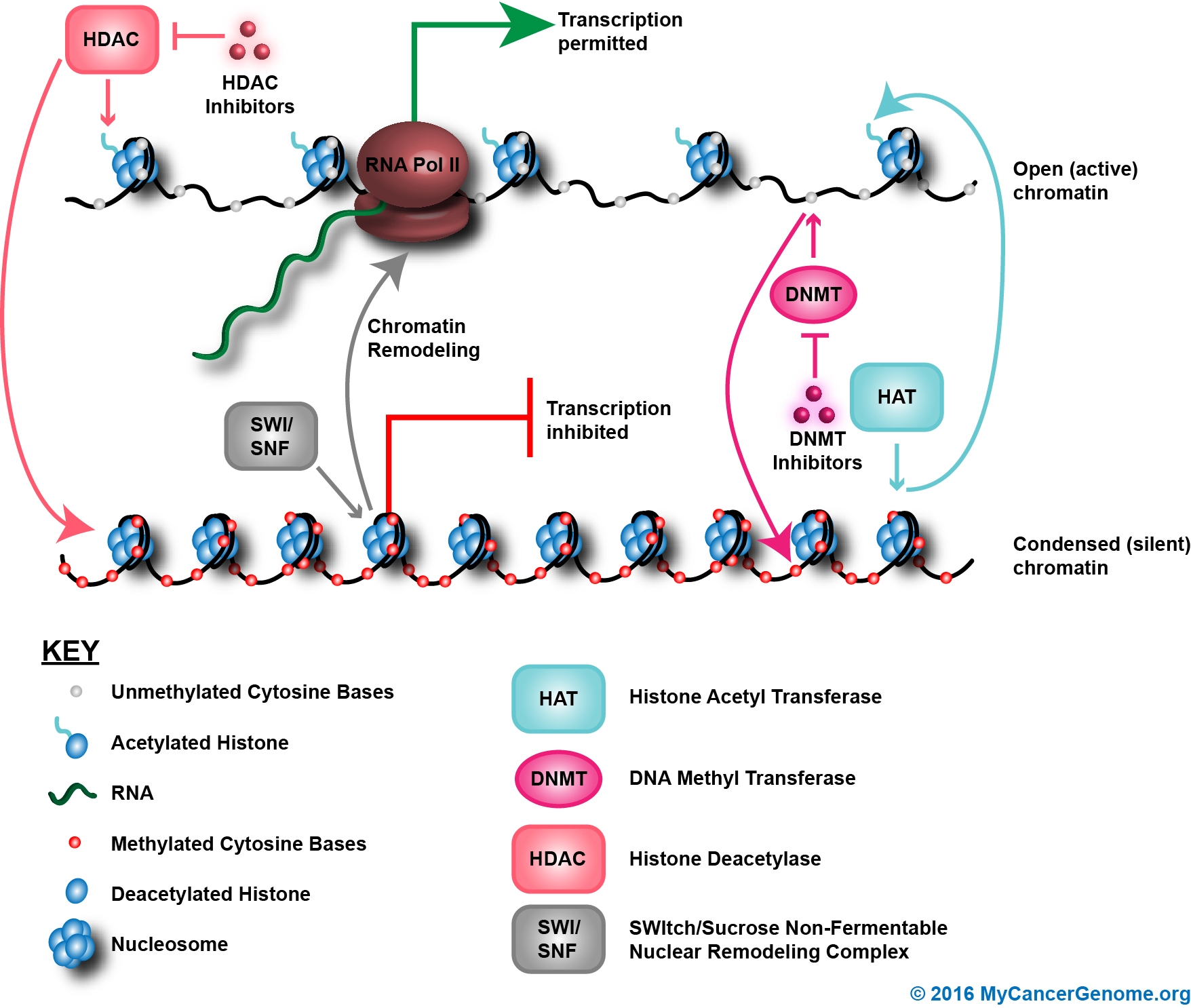
Figure 1. DNA methylation is an epigenetic process of chromatin remodeling that regulates gene expression. Methylation of cytosine residues by DNA methyltransferase represses transcription and switches genes off. The addition of acetyl groups to histones by histone acetylase activates transcription and switches gene on. Specific nodes in the pathway that are therapeutically actionable are noted.
 Back to Pathways List
Back to Pathways List